R Studio crash with spacyR, what now ?
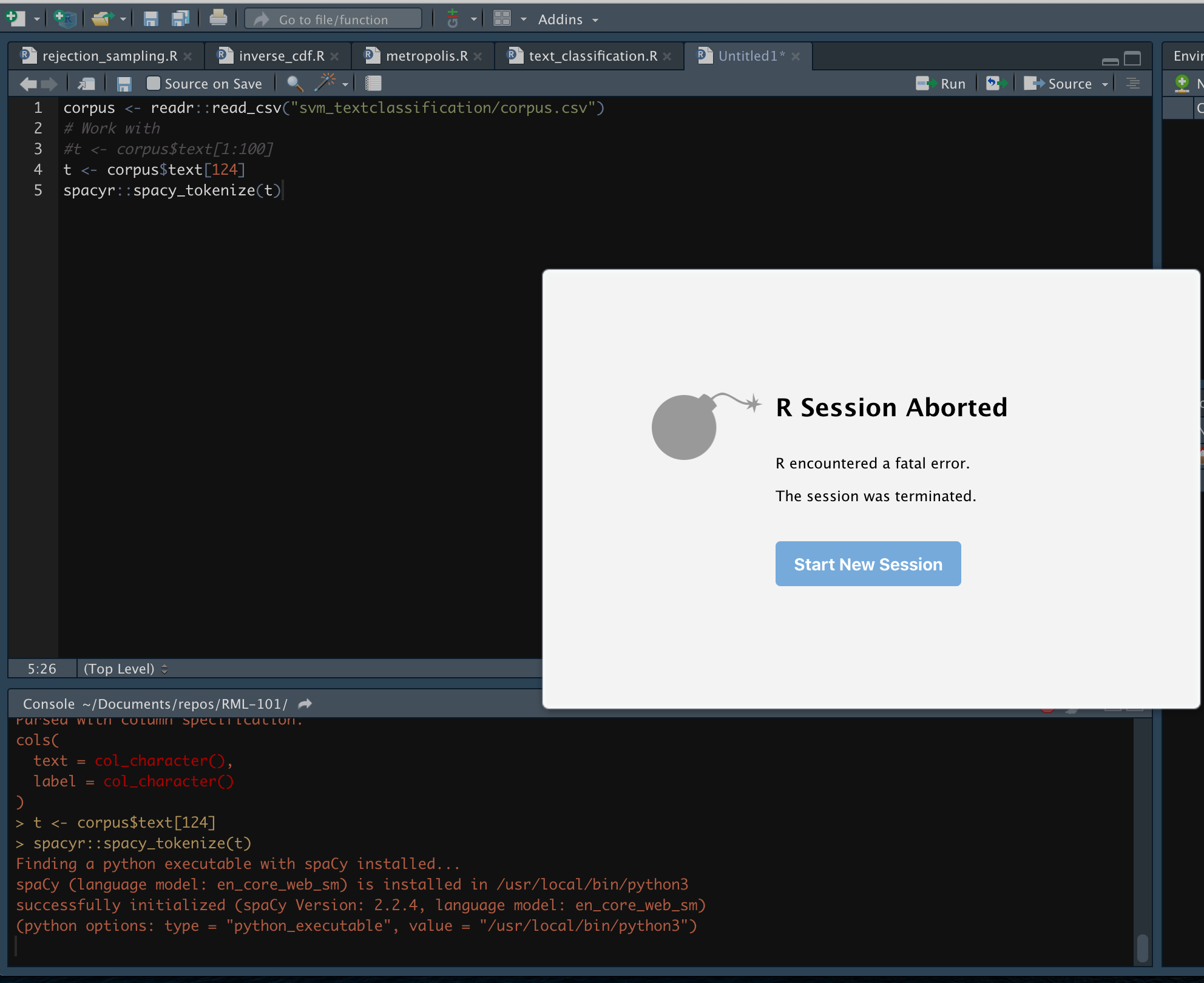
I recently posted an example on Text classification modelling with tidyverse, SVM vs Naivebayes, as usual when one is working with data unexpected things can happen, in this particular case it was a rather strange crash in RStudio that took me a couple of hours to figure out. TBH I think I spent as much time looking at this issue than the code for the post, but I digress. Let’s take a look at the isolated code that made R Studio kaboom:
corpus <- readr::read_csv("svm_textclassification/corpus.csv")
# Work with t <- corpus$text[1:100]
t <- corpus$text[124]
spacyr::spacy_tokenize(t)As you can see the issue appear to be with spacyr package, in fact I was so conviced that it was a bug that I even submitted an official issue to their github repository, spacyr::spacy_tokenize(t) crashes R address 0x8, cause ‘memory not mapped’ with specific string . The main issue seems to be with the spacy_tokenize function but the problem is that there was no feedback from the IDE.
There is an excellent guide for Debugging with RStudio but I couldnt find an scenario to which the IDE itself crash. So what now ?
Using R command
In order to skip the issue that might have arise within the IDE, you can go straight to using R in the command line, and run your script from there. This is particularly useful because any issue that was happening was probably outside the scope of R itself, and since spacyR make heavy use of reticulate in order to access spacy library function in python this was a good starting point. Then once the code above is run through the console we get:
*** caught segfault ***
address 0x8, cause 'memory not mapped'
Traceback:
1: r_to_py_impl(x, convert = convert)
2: r_to_py.default(values)
3: reticulate::r_to_py(values)
4: eval(parse(text = sprintf("main$%s <- reticulate::r_to_py(values)", pyvarname)))
5: eval(parse(text = sprintf("main$%s <- reticulate::r_to_py(values)", pyvarname)))
6: spacyr_pyassign("texts", x)
7: spacy_tokenize.character(t)
8: spacyr::spacy_tokenize(t)
Possible actions:
1: abort (with core dump, if enabled)
2: normal R exit
3: exit R without saving workspace
4: exit R saving workspace
Selection:
Et voila! There it’s the error when calling a function r_to_py_impl(x, convert = convert) this is a great clue meaning that the problem is outside R itself. But what is it ? Well it turns out that the issue was the encoding I was using, assuming that the corpus.csv was encoded with standard UTF-8 instead of ISO-8831 for latin characters. Therefore the solution was rather simple
corpus <- readr::read_csv("svm_textclassification/corpus.csv", locale = locale(encoding = "latin1"))
# Work with
#t <- corpus$text[1:100]
t <- corpus$text[124]
spacyr::spacy_tokenize(t)So remember anytime you encounter this type of crashing issue, go straight to the R command in the console line to get more information about what is going on.